How Do You Know if Your Transformation Was Successful
1.13: Transformation
- Page ID
- 36755
Learning Objectives
Goals:
- Explicate how the information encoded in a gene is expressed as a trait
- Describe the role of transformation in cloning genes
- Explain the purpose of each control in the transformation experiment
Student Learning Outcomes:
Upon completion of this lab, students volition exist able to:
- Carryout a transformation
- Predict the growth results for the negative control and plasmid containing reaction on both antibody-containing and nutrient agar only media
- Explain your reasoning, if predicted growth results don't match actual growth results
Introduction
Genetic engineering or DNA technology has been useful for producing large quantities of a specific protein to treat human being diseases. For case, patients with diabetes, hemophilia, or anemia require treatments with insulin, clotting factor, and growth factor proteins. Targeted genes (DNA) can exist cut with restriction enzymes and joined with other Deoxyribonucleic acid with the enzyme ligase. A cloning vector is used to carry the recombinant DNA into living cells, and then that the cells can synthesize the encoded proteins. The all-time cloning vectors are pocket-size in size, able to replicate its DNA, incorporate restriction enzyme recognition sites, and have a marker gene (ordinarily antibiotic resistance gene). In this lab, we will employ a recombinant plasmid as the cloning vector. This recombinant plasmid contains (1) a promoter that enables transcription of desired factor, (two) a sequence for the initiation of Deoxyribonucleic acid replication (ori site), and (3) an antibiotic resistance cistron.
Transforming Bacteria with Recombinant Plasmid
Inserting a gene into a plasmid vector is an important first step in the gene cloning process. Still, if the ultimate goal is to produce a large amount of a particular protein, the plasmid must replicate to make sure that there are many copies of the gene and the gene of interest must be expressed, meaning the gene is utilized to produce the encoded protein. Both activities can only occur inside a cell. Therefore, in this lab we will put a recombinant plasmid into Due east. coli bacteria through a process that is called transformation, and then named because it changes the Dna content of the leaner.
The plasmid will be taken upward by bacteria where it replicates, and its genes volition be expressed using the bacterial cellular mechanism. If a gene of interest has been inserted into the plasmid vector, the bacteria produces the product encoded by that gene.
In this exercise, you volition carry out the transformation of E. coli leaner using a recombinant plasmid that contains a gene that produces colored proteins.
Bacterial Transformation
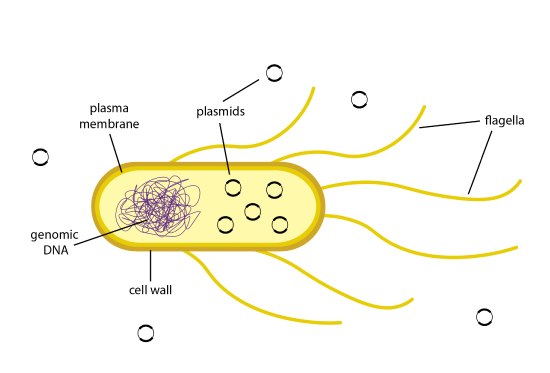
Once a recombinant plasmid is fabricated that contains a gene of involvement, such every bit insulin, the plasmid can enter bacterial cells by a process called transformation. Effigy 13.1 illustrates transformation. The uptake of Dna from the surround of a bacterial cell occurs with a very low efficiency in nature. Due east. coli bacteria take complex plasma membranes that divide the external environment from the internal environment of the cell and advisedly regulate which substances can enter and exit the jail cell. In addition, the cell wall is negatively charged and repels negatively charged Deoxyribonucleic acid molecules.
Cells that have been treated to become competent are more efficient at taking in DNA from their surrounding environment. Competent cells tin can be made by treating the bacteria with a calcium solution. Calcium ions are positively charged, and will neutralize the negatively charged outer membrane on the East. coli bacteria. With the positive accuse now coating the membrane, the inherently negatively charged Deoxyribonucleic acid molecules will move through the plasma membranes and into the cell. The transformation efficiency can be further increased by stressing the cells in a oestrus stupor. By changing the temperature of the cells drastically from cold to warm, the plasma membranes go more fluid and create pores in them. The plasmid Deoxyribonucleic acid can travel from the surround through these pores and enter the prison cell. The cells are then plunged back into a cold temperature, which causes the pores to shut and the plasmid Deoxyribonucleic acid to remain inside the cell.
Nevertheless, even competent cells exercise non always uptake the plasmid. For some plasmid DNA molecules, only near ane in 10,000 cells volition be transformed. When so few cells accept taken in the plasmid, how volition you be able to identify transformed cells? When designing a recombinant plasmid, one of the requirements is to add together a gene for an antibody resistance. This way, the bacteria tin can be grown in the media with an antibiotic added to it, and only cells that take the resistance gene, such as those that express the recombinant plasmid, will be able to grow.
From Plasmid DNA to Protein
After a recombinant plasmid enters a bacterial cell, the cell begins to limited the genes on it. Dna polymerase locates the ori- the origin of replication, and starts to replicate the plasmid using the bacterial cell'due south machinery. Multiple copies of the recombinant plasmid tin enable the bacterial cell to express large amounts of a protein. Unremarkably, a bacterial prison cell will only make the protein of involvement, afterward it is induced to exercise so by adding a chemical which will promote the transcription of the gene. Retrieve that to limited the cistron encoding the protein on the recombinant plasmid, DNA is transcribed to mRNA, which is then translated to protein (Effigy xiii.two). The expressed proteins may touch on the visible traits when observing the bacteria colonies.
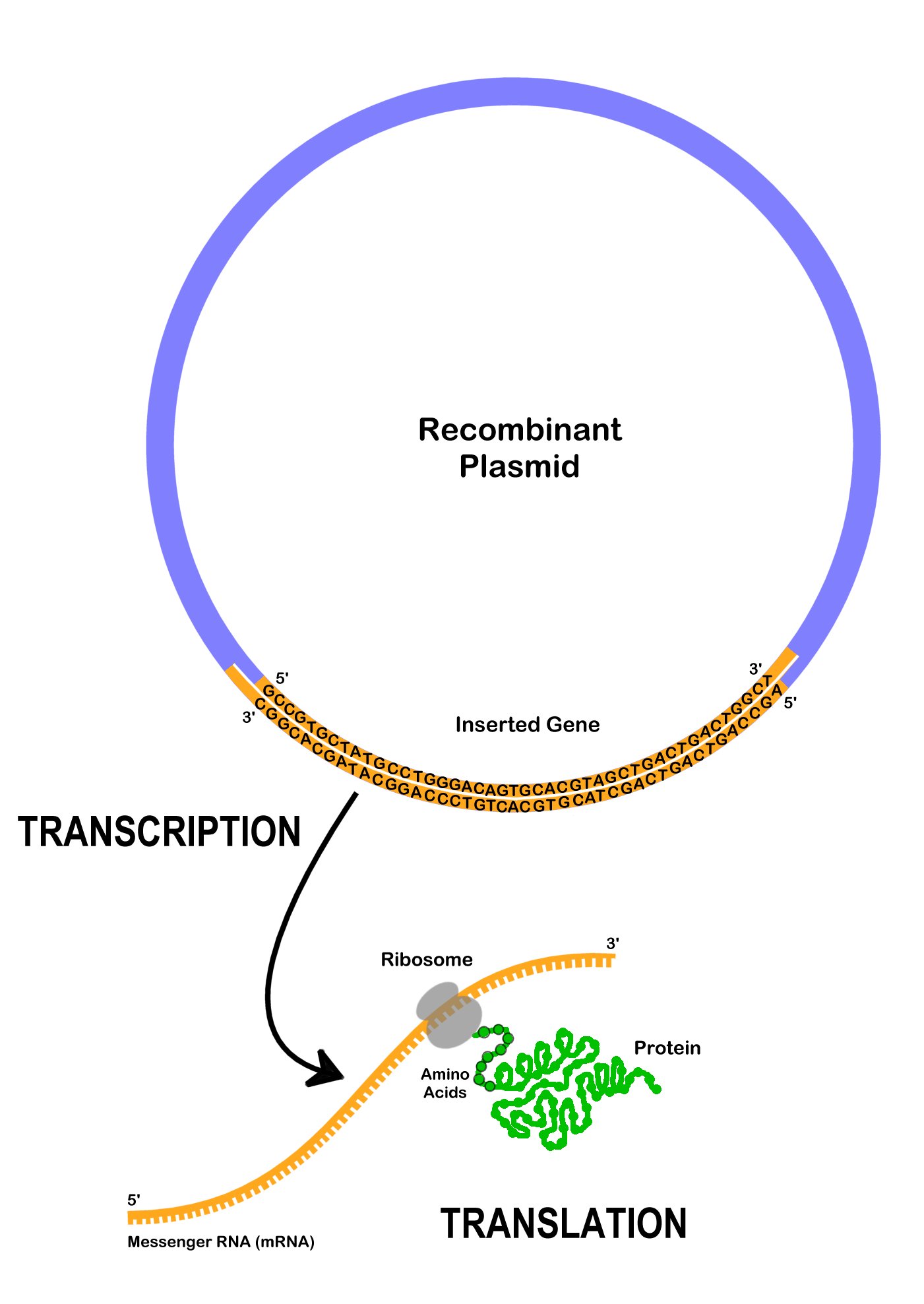
Recombinant plasmids and other forms of genetic engineering is possible because all living organisms use DNA every bit a platform to encode genetic information. Genes from unlike organisms can be expressed in other organisms like bacteria since they are encoded in DNA. The Deoxyribonucleic acid instructions can be transferred, and other organisms can limited foreign traits.
Proteins have many different functions inside and cells. They are made upward of smaller subunits, amino acids, which are encoded past DNA nucleotides. A specific 3 nucleotide sequence that codes for a single amino acrid is called a codon. For case, the codon TTG codes for the amino acid tryptophan, whereas the codon AAG codes for the amino acid lysine. In many cases, more than than one codon can encode the same amino acid. For example, AAA is also a codon for lysine. In addition, there are informational codons, such as the starting time codon (ATG) and the stop codon (TTA), which show where in the Dna sequence the code for the protein begins and ends.
Transforming Leaner with Plasmids
In this laboratory experiment you will transform E. coli leaner cells with plasmids. You will be using E. coli that has been made competent with a calcium chloride treatment, and form two dissimilar testing groups: a negative control cell group that does not have plasmids added to it, and the experimental grouping that has the plasmids added. Later on the cells are heat-shocked, they will be grown under various testing conditions:
- The control group on nutrient agar (a type of growth media that bacteria thrive on).
- The control group on nutrient agar with an antibiotic added.
- The experimental grouping on nutrient agar.
- The experimental group on nutrient agar with an antibiotic.
- The experimental group on food agar, antibody and an inducer (such every bit IPTG).
By examining the growth of bacteria under these conditions, you can verify that your procedure worked, and y'all can place the bacteria transformed with the added plasmid. How will you know if y'all are successful? In the examples for plasmids we have recommended for this exercise, the recombinant bacteria will have a new and highly visible trait: It will now produce colored protein, which makes the cells themselves colored! As the bacteria multiply on the media, they form visible collections of cells called colonies. Each colony represents the decedents of the original bacterial cell that landed on that spot on the medium and began to replicate. Thus colonies are clones (verbal copies) of the prison cell that began the replication process.
The relevant components of your plasmid are the gene for the colored protein, the inducible promoter, and the ampicillin resistance factor (ampR). The ampR factor confers resistance to the antibiotic ampicillin. (Biotechnologists phone call these genes selectable markers because just bacteria having the gene will survive in the presence of an antibiotic.) If the inducer is present in the bacteria, the promoter will exist "turned on" and so RNA polymerase can transcribe the gene of involvement. This volition allow poly peptide to be produced.
Prelab Questions
Discuss the following among yourselves. Be ready to share your thoughts with the rest of the class.
- Ampicillin is a derivative of the antibiotic penicilliin. It disrupts cell wall formation in bacterial cells which kills the cells. Nonetheless, our recombinant plasmid contains a cistron that provides antibiotic resistance past producing a protein that breaks downwardly ampicillin. Why do we include ampicillin in the exam medium?
- What will happen if the transformed cells do not grow in the presence of the chemic inducer?
- In the experiment, yous will add the command and experimental groups of cells onto different media combinations. What exercise you predict for each status? Fill in Table 1 by indicating if you predict growth or no growth, and if growth, volition there exist minimal growth or lots of growth.
Read through the Procedures below and outline the steps, using words and a flowchart in your lab notebook.
| Medium | No plasmid control | Treatment with plasmid |
|---|---|---|
| Nutrient Agar | ||
| Nutrient Agar + Ampicillin | ||
| Nutrient Agar + Ampicillin + Inducer |
Transforming E. coli
MATERIALS
Reagents
- Plasmid – in microfuge tube
- Nutrient Goop (NB) – in microfuge tube
- Competent E. coli cells (CC) – in microfuge tube (Always go along CC tube on ice)
- 3 agar plates:
Plate 1: NA
Plate 2: NA/amp
Plate iii: NA/amp/ind
Supplies and Equipment
- P-20 micropipette
- P-200 micropipette
- Pipette tip box (for P-20, P-200)
- Microfuge tube rack
- Two 1.5 mL microfuge tubes
- Permanent marker
- Disposable gloves
- Crushed water ice in a Styrofoam loving cup (fill cup outset with ice before taking CC tube.)
- Pack of jail cell spreaders (do not remove spreaders from pack until directed to do so)
- Timer or clock
- Floating microfuge tube rack
- 42°C h2o bathroom
- 37°C incubator
- Tape
- Waste container
- Biohazard bag (for supplies that handle cells)
Rubber
Check your protocol and follow all safety measures and wearable proper attire prior to conducting the experiment.
Exercise hygienic technique while using E. coli or other live specimens in a laboratory setting. Aseptic technique is the practice of taking precautions to limit potential contagion to both the person performing the experiment, and to the sample/due south. Please annotation the following:
- Wear gloves when working with bacteria.
- Avoid touching contaminated areas which includes annihilation that has touched bacteria. Notify your instructor ASAP if an accident takes place, such as a spill.
- Put all supplies that accept been exposed to leaner into either a biohazard bag, or a designated biowaste container. These contaminated supplies may include pipette tips, prison cell spreaders, and microfuge tubes.
- Proceed agar plates closed at all times afterwards removing them from the incubator.
- Always launder your hands for 20 seconds with soap and h2o before leaving the lab.
Process
- Make sure yous have all the reagents in a tube rack.
- Retrieve a chilled CC tube and put in the cup of crushed ice. Continue the competent cells cold at all times. Hold the tube past the rim, not the bottom.
- Characterization the acme and sides of 2 new microfuge tubes with "P-" and "P+".

- Put the P– and P+ tubes with the CC tube on ice.
To ensure the all-time results possible, it is crucial that each step is followed exactly. Limit whatever possible contamination to the materials, yourself, and environs. - Add E. coli competent cells (CC tube) to both the P- and P+ tubes.
- Take the P-200 pipette, set to l µL and put on a tip.
- Holding the CC tube by the rim, gently resuspend the cells by slowly pumping between the first and no terminate with the pipette (a gentle downward plunger motion to the beginning resistance point then a gentle up motion to the top plunger position).
- Add 50 µL of cells to the P+ tube and place the P+ tube on ice immediately. Discard the tip into the sharps container.
- Grab a new pipette tip. Repeat for the P- tube. Discard the tip. Both P- and P+ tubes should be on ice and contain 50 µL of competent cells.
- Add plasmid to the P+ tube only.
- Take the P-twenty pipette, set to x µL and put on a tip.
- Remove 10 µL of the plasmid and add it to the P+ tube. Mix together by slowly pumping between the kickoff and no cease ii-3 times then place the tube back on ice.
- Put the P- and P+ tubes on water ice for xv minutes.
- While the tubes are chilling, obtain your agar plates and marker. Exercise non open the plates during this step
- Each agar plate contains different media- one of Nutrient Agar only (NA), one of Nutrient Agar + ampicillin (NA/AMP), and ane of Food Agar + ampicillin + inducer (NA/AMP/IND). These may be labeled with a stripe pattern or written on the plate
- Exercise not open the plates. Turn every plate upside-downward; the agar should be on height. Label the agar side with 1) appointment 2) grouping number three) class menstruation. Endeavor to write pocket-sized along the lesser edge of the plate.
- Next, draw a line downwardly the eye of the NA and NA/AMP plates. I half is labeled as "P-" and the other is "P+". The NA/AMP/IND is simply labeled "P+". They should expect like to Fig 4
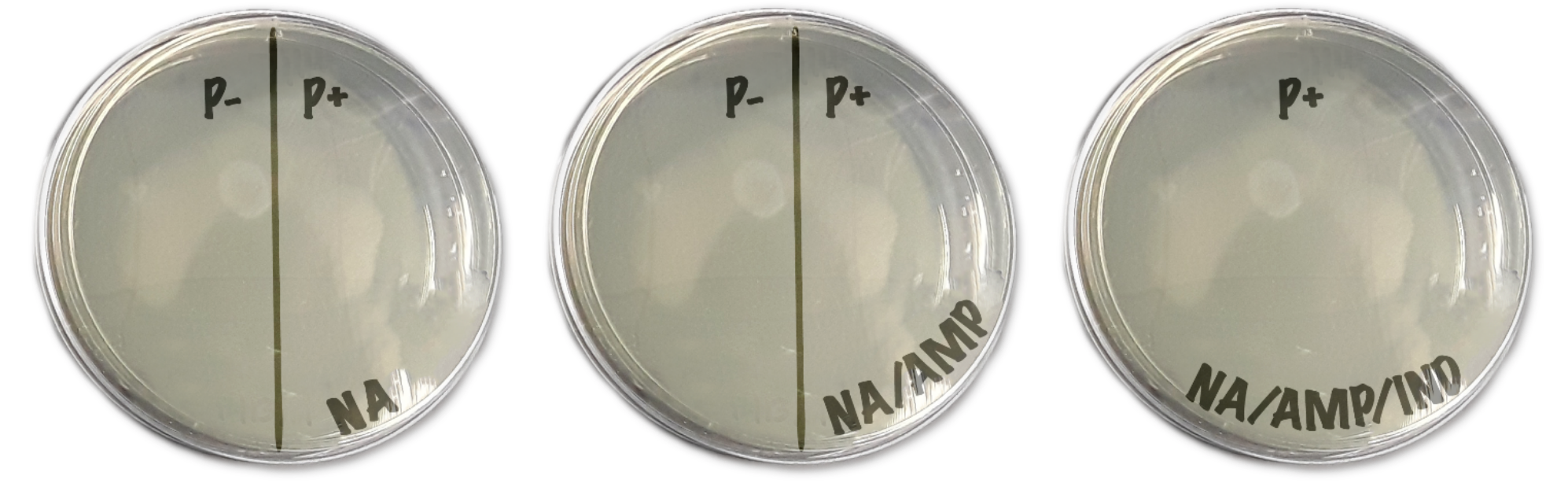
- After the P- and P+ tubes have been on water ice for 15 minutes, keep the tubes on ice and bring the water ice cup over to the 42°C water bathroom. Yous will also need your timer/clock. Put both tubes into a floating microfuge tube rack, then identify information technology into the water bathroom for precisely 45 seconds.
- As soon every bit the 45 seconds pass, immediately put the tubes back into the ice loving cup and proceed on ice for a minimum of 1 minute.
- Add Nutrient Broth (NB) to the P- and P+ tubes.
- Take the P-200 pipette, set to 150 µL and put on a tip.
- Remove 150 µL of NB and add to the P- tube. Mix together by slowly pumping between the first and no stop with the pipette. Discard the tip into the biowaste container.
- Get a new pipette tip. Echo the same procedure for the P+ tube.
- Keep the tubes at room temperature for xv minutes. If you are running short on time, this step can be shortened.
- Add cells from the P– tube onto your NA and NA/amp plates. Keep the plates right side upward and so the agar is on the bottom. You add cells to the surface of the agar (non the plastic lid).
- Take the P-200 pipette, gear up to 50 µL and put on a tip
- Have the P- tube. Slowly pump the pipette between the first and no stop with the pipette to resuspend the cells. Remove 50 µL of the cells.
- Lift the hat of the NA plate like a "clamshell" to go out a large enough gap to deliver the cells, while lowering the chance of airborne contamination. Add the 50 µL of cells to the P- half of the plate. Shut the plate and fix to spread the cells.
- Repeat this for the NA/AMP plate by resuspending the P- tube cells with the pipette and aforementioned tip. Add 50 µL of the cells to P- side of the NA/AMP plate using the clamshell method again. Discard the tip into the sharps container.
- Spread the cells from the P– tube on your NA and NA/amp plates. You must spread your plates in this lodge.
- Open up the sterilized cell spreader package. Take a single spreader out and concord it merely by the end you removed it by. Be careful not to touch the other cease of the spreader to anything only the cells and agar. Shut the packet.
- Using the "clamshell" method, open the NA lid and spread the cells on the P- one-half of the plate. Gently concur the spreader flat against the surface of the agar; treat it gently every bit if you were handling gelatin. Close the plate.
- Repeat the same technique for the NA/AMP plate on the P- side using the same spreader. Once you are finished, discard the spreader into the designated biowaste container.
- Add cells from the P+ tube to your NA, NA/amp, and NA/amp/ind plates:
- Take the P-200 pipette, double check it is set to fifty µL and put on a tip.
- Accept the P+ tube. Slowly pump the pipette between the first and no stop with the pipette to resuspend the cells. Remove 50 µL of the cells.
- Open the NA plate again like a clamshell to evangelize 50 µL of cells to the P+ half of the plate. Close the plate.
- Repeat this for the NA/AMP plate by resuspending the cells with the pipette and aforementioned tip. Add 50 µL of cells to the P+ half of the plate. Close the plate.
- Repeat this for the NA/AMP/IND plate by resuspending the cells with the pipette and same tip. Add together l µL of cells ii times to the entire plate. There is a total of 100 µL of P+ cells added to the plate and must cover the entire surface when spread. Close the plate.
- Spread the P+ cells on the NA, NA/AMP and NA/AMP/IND plates. You must spread your plates in this social club.
- Open the cell spreader bundle again and take out a single spreader, simply touching the handle. Practise Be conscientious to not touch the other finish of the spreader to anything but the cells and agar.
- Using the clamshell method, open the NA lid and spread the cells on the P+ half of the plate. Gently hold the spreader flat against the surface of the agar; treat it gently as if you were handling gelatin. Close the plate.
- Using the same spreader, repeat the same technique for the NA/AMP plate on the P+ side .
- Repeat the aforementioned technique for the NA/AMP/IND plate except the cells must be spread across the entire plate, not just i half. Rotate the plate gently to evenly disperse the cells with the spreader. In one case you are finished, discard the spreader into the designated biowaste container.
- Proceed all plates right side up for five minutes until the liquid is fully captivated into the plate. Stack the plates together and tape them, labeling the tape with your class period, grouping number and date.
- Put the plates upside down (agar side on meridian) into the 37°C incubator to forbid condensation from forming and falling on the cells.
- Put anything that touched the cells into the biohazard handbag, including pipette tips, microfuge tubes and cell spreaders. Wipe down tabletops with disinfectants and wash easily.
- Incubate the plates at 37°C for 24-36 hours and await for growth. Go along the plates closed.
- Put the agar plates into the biohazard bag when told to do so.
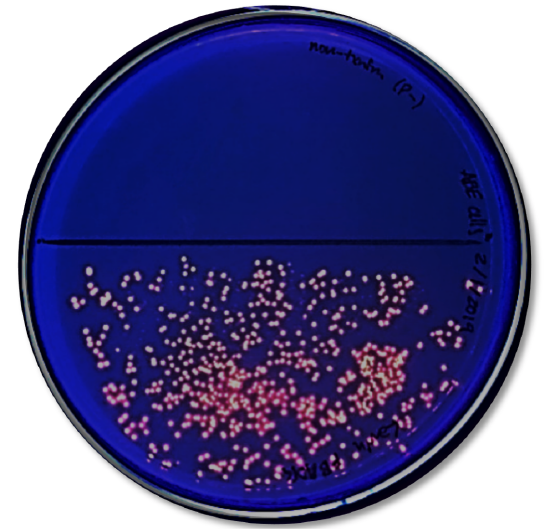
Analysis
- Look at the results of your transformation. Fill out Table two with observations on whether you meet growth or not on the unlike media.
- Do your actual results lucifer your predicted results? If not, what differences do you see and what are some explanations for these differences?
- How many colored colonies were present on your NA/AMP/IND plate?
| Medium | No plasmid command | Treatment with plasmid |
|---|---|---|
| Nutrient Agar | ||
| Food Agar + Ampicillin | ||
| Nutrient Agar + Ampicillin + Inducer |
Study Questions
- Why would colored colonies form on the NA/AMP/IND plate and not the NA/AMP plate?
- What are some possible reasons for colored colonies to form on the NA/AMP plate?
- Extrachromosomal Deoxyribonucleic acid in bacteria, similar a recombinant plasmid, tin can replicate within the cell without the balance of the prison cell'due south DNA replicating. This tin atomic number 82 to multiple plasmids within a cell. Why is this important?
- Previously, you learned about the interactions between Dna, RNA, proteins and traits. Explain how an inserted gene such as 1 in plasmid Deoxyribonucleic acid, is expressed as a trait.
- How can bacteria make proteins from strange Deoxyribonucleic acid, such as human insulin or a protein from a jellyfish like green fluorescent protein (GFP)?
- What would happen if we grew the transformed bacteria in the presence of a unlike antibiotic such equally kanamycin, instead of ampicillin?
barhamramessanies89.blogspot.com
Source: https://bio.libretexts.org/Bookshelves/Biotechnology/Lab_Manual%3A_Introduction_to_Biotechnology/01%3A_Techniques/1.13%3A_Transformation

Post a Comment for "How Do You Know if Your Transformation Was Successful"